Specialized Antibody Characterization
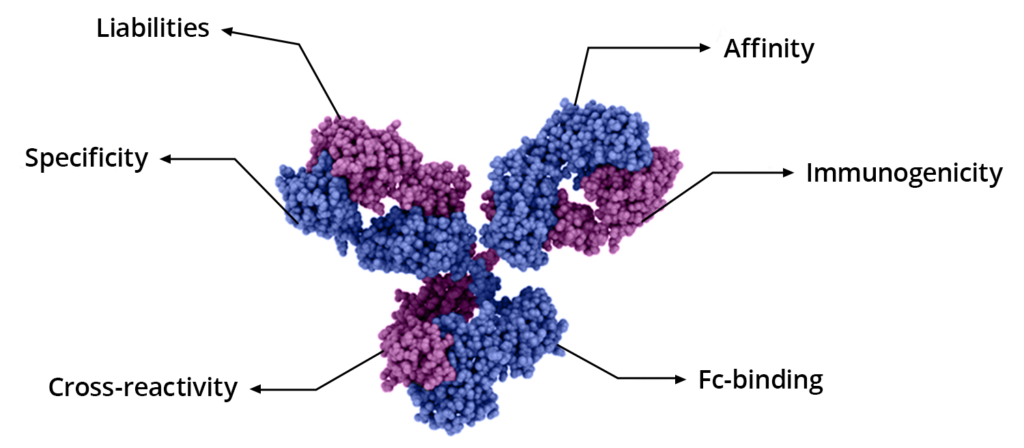
In addition to discovering your next target-binding antibody for diagnostic or therapeutic applications, additional characterization such as affinity analysis, epitope binning, FcR binding, and in silico predictions (immunogenicity analysis and sequence liability) can be achieved using an assortment of methods.
Affinity Analysis
We employ two very powerful platforms to determine the kinetic properties of antibody-antigen interactions of test antibodies or antibody-fragments as either purified or non-purified samples:
- High-throughput array SPR by Carterra LSA (as high as 384 antibodies tested per plate)
- BLI-based Octet system
Epitope Binning
Epitope binning investigates the sequential binding of two antibodies to an antigen. Antibodies with similar blocking profiles are grouped into bins specific to the same or overlapping epitopes. Curia uses two proven platforms to test the ability of multiple antibodies to block each other’s binding to the antigen.
- High-throughput epitope binning by arrayed SPR using the Carterra LSA (as high as 96 x 96 antibodies tested per plate)
- BLI platform-based Octet system (as high as 16 x 16 antibodies tested)
Curia is an authorized partner of Carterra® and the only CRDMO with a Carterra® LSA™ instrument.
FcRy and FcRn Binding Assays
Measuring an antibody’s affinity to the Fcγ receptors can be critical for predicting biological and pharmacological outcomes, such as cytotoxicity or half-life duration. For your biotherapeutics development, Curia has established a standard human Fcγ receptor panel binding assay in a high-throughput format using the BLI platform. This in vitro binding assay can also be used for quality control purposes in parallel with functional cell-based ADCC. Additionally, Curia offers clients an FcRn binding assay to study pH-dependent interaction of antibody with the neonatal Fc-receptor (FcRn) in high-throughput and easy to use format. The outcome of the study will enhance overall understanding of the antibody and demonstrate comparability in the development of biosimilars.
Sequence Liability Identification
Post-translational modifications such as glycosylation, deamidation, isomerization, and oxidation have the potential to reduce affinity, potency, stability, and homogeneity of an antibody, resulting in complications in downstream process development. Curia has developed algorithms to identify sequences within antibody variable regions that could present potential liabilities to product quality. The sequence liability identification service can help the investigator to assess whether they need to be eliminated early in the engineering process and to select liability free antibodies for further development.
In Silico Immunogenicity Prediction
Biotherapeutic antibodies and proteins have the risk of causing anti-drug antibody responses in human bodies and fail to achieve desired therapeutic goals or cause allergic reactions. Major histocompatibility complex class II (MHC class II) are the key molecules to recognize the immunogenic peptides, presented as epitopes from the biotherapeutic molecules, on antigen presenting cells and imprint the T cell receptors for cascade responses. Our in silico immunogenicity prediction service provides the cost-effective initial analysis of the immunogenicity of the biologics against the major MHC class II alleles. Using trained artificial neural network ensembles, nine core residues with potential affinity to the binding cleft of MHC class II molecules are identified, providing important information for further engineering and optimization.
| Catalog # | Name | Timeline | Price |
|---|---|---|---|
| 1018 | Epitope Binning | 2 to 4 weeks | Request |
| 24160 | Affinity Measurement | 2 to 3 weeks | Request |
| 24190 | Fcγ receptor panel binding assay | 1 to 2 Weeks | Request |
| 24700 | Sequence Liability Identification | 4 to 5 Days | Request |
| 24701 | In Silico Immunogenicity Prediction | 4 to 5 Days | Request |
Antibody Discovery & Engineering Services
The Curia Antibody Center offers comprehensive antibody discovery and engineering services, including identification, optimization, and characterization of new leads.
